Smart Citations
Smart CitationsSee how this article has been cited at scite.ai
scite shows how a scientific paper has been cited by providing the context of the citation, a classification describing whether it supports, mentions, or contrasts the cited claim, and a label indicating in which section the citation was made.
Immunohistochemical markers of CYP3A4 and CYP3A7: a new tool towards personalized pharmacotherapy of hepatocellular carcinoma
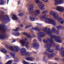
Hepatocellular carcinoma (HCC) represents a major global health problem, since more than 90% of primary liver cancers worldwide are HCC. Most cases of HCC are secondary to viral hepatitis infection (hepatitis B or C), alcoholism and cirrhosis. Sorafenib, an oral tyrosine kinase inhibitor that suppresses tumor proliferation and angiogenesis, emerged as the first effective systemic treatment for HCC after 30 years of research, and is currently the standard-of-care for patients with advanced HCC. Sorafenib is metabolized by cytochrome P450 (CYP450), particularly from the 3A4 isoform, producing two main metabolites: the N-oxide and the N-hydroxymethyl metabolite. We studied 11 HCC sample showing the presence of CYP3A4 and CYP3A7 in most of the samples analyzed. Specifically, the immunoreactivity of CYP3A4 was more strong and widespread than that of CYP3A7. The CYP3A4 immunoreactivity was observed in surrounding hepatocytes in 8 out of 11 cases; while the CYP3A7 immunostaining was found in normal liver cells, in 7 out of 11 cases. These results suggest the existence of a marked inter-individual variability regarding the presence of the isoforms of CYP3A. In addition, since sorafenib is metabolized by CYP3A4, but not by CYP3A7, an overexpression of CYP3A4 may lead to an increase in the degradation of the drug and then to clinical ineffectiveness. These results might implicate the necessity of an individualized approach in the treatment of HCC as positivity to CYP3A4 in HCC liver samples might predict a scarce response to sorafenib.
How to Cite
PAGEPress has chosen to apply the Creative Commons Attribution NonCommercial 4.0 International License (CC BY-NC 4.0) to all manuscripts to be published.