Smart Citations
Smart CitationsSee how this article has been cited at scite.ai
scite shows how a scientific paper has been cited by providing the context of the citation, a classification describing whether it supports, mentions, or contrasts the cited claim, and a label indicating in which section the citation was made.
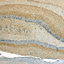
Expression of the endocannabinoid receptors in human fascial tissue
Cannabinoid receptors have been localized in the central and peripheral nervous system as well as on cells of the immune system, but recent studies on animal tissue gave evidence for the presence of cannabinoid receptors in different types of tissues. Their presence was supposed also in myofascial tissue, suggesting that the endocannabinoid system may help resolve myofascial trigger points and relieve symptoms of fibromyalgia. However, until now the expression of CB1 (cannabinoid receptor 1) and CB2 (cannabinoid receptor 2) in fasciae has not yet been established. Small samples of fascia were collected from volunteers patients during orthopedic surgery. For each sample were done a cell isolation, immunohistochemical investigation (CB1 and CB2 antibodies) and real time RT-PCR to detect the expression of CB1 and CB2. Both cannabinoid receptors are expressed in human fascia and in human fascial fibroblasts culture cells, although to a lesser extent than the control gene. We can assume that the expression of mRNA and protein of CB1 and CB2 receptors in fascial tissue are concentrated into the fibroblasts. This is the first demonstration that the fibroblasts of the muscular fasciae express CB1 and CB2. The presence of these receptors could help to provide a description of cannabinoid receptors distribution and to better explain the role of fasciae as pain generator and the efficacy of some fascial treatments. Indeed the endocannabinoid receptors of fascial fibroblasts can contribute to modulate the fascial fibrosis and inflammation.
Downloads
Publication Facts
Reviewer profiles N/A
Author statements
- Academic society
- N/A
- Publisher
- PAGEPress Publications, Pavia, Italy
Citations
10.1515/jcim-2020-0013
10.4081/ejh.2022.3426
10.1210/jc.2019-00665
10.1016/B978-2-294-76430-1.00009-3
10.56986/pim.2024.10.002
10.1007/s00337-016-0172-1
10.12677/ACM.2021.114228
10.3389/fmed.2024.1472116
10.3390/ijms21082936
10.2478/acb-2023-0002
10.1007/s11033-022-07223-5
10.1371/journal.pone.0181064
How to Cite
PAGEPress has chosen to apply the Creative Commons Attribution NonCommercial 4.0 International License (CC BY-NC 4.0) to all manuscripts to be published.

 https://doi.org/10.4081/ejh.2016.2643
https://doi.org/10.4081/ejh.2016.2643