Smart Citations
Smart CitationsSee how this article has been cited at scite.ai
scite shows how a scientific paper has been cited by providing the context of the citation, a classification describing whether it supports, mentions, or contrasts the cited claim, and a label indicating in which section the citation was made.
The possible prognostic role of histone deacetylase and transforming growth factor β/Smad signaling in high grade gliomas treated by radio-chemotherapy: a preliminary immunohistochemical study
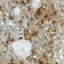
Glioblastoma (GBM) is the most common and aggressive tumor of the central nervous system. Unfortunately, patients affected by this disease have a very poor prognosis, due to high level of invasiveness and resistance to standard therapies. Although the molecular profile of GBM has been extensively investigated, the events responsible for its pathogenesis and progression remain largely unknown. Histone Deacetylases (HDAC) dependent epigenetic modifications and transforming growth factor (TGF)-β/Smad pathway seem to play an important role in GBM tumorigenesis, resistance to common therapies and poor clinical outcome. The aim of this study was to evaluate the involvement and the possible interaction between these two molecular cascades in the pathogenesis and prognosis of GBM. Immunohistochemistry (IHC) was performed on microdissected GBM samples, collected from 14 patients (6 men and 8 women) ranging in age from 43 to 74 years. The patients were previously divided, on the basis of their overall survival (OS), into two groups: short and long OS. Patients with poor prognosis showed hyperexpression of HDAC4 and HDAC6, an activation of the TGF-β/Smad pathway, with high levels of IL-13, Smad2, PDGF and MMP3 expression, compared to the long survivors. The short OS group exhibits a decrease in Smad 7 expression and also low levels of p21 immunostaining, which represents a common target of the two pathways. The IHC data was confirmed by quantitative analysis and Immunoblotting. Our preliminary results suggest that both HDAC4 and HDAC6 together with the TGF-β/Smad pathway may be involved in progression of GBM and this cross talking could be a useful prognostic marker in this deadly disease.
Downloads
Publication Facts
Reviewer profiles N/A
Author statements
- Academic society
- N/A
- Publisher
- PAGEPress Publications, Pavia, Italy
How to Cite
PAGEPress has chosen to apply the Creative Commons Attribution NonCommercial 4.0 International License (CC BY-NC 4.0) to all manuscripts to be published.

 https://doi.org/10.4081/ejh.2017.2732
https://doi.org/10.4081/ejh.2017.2732