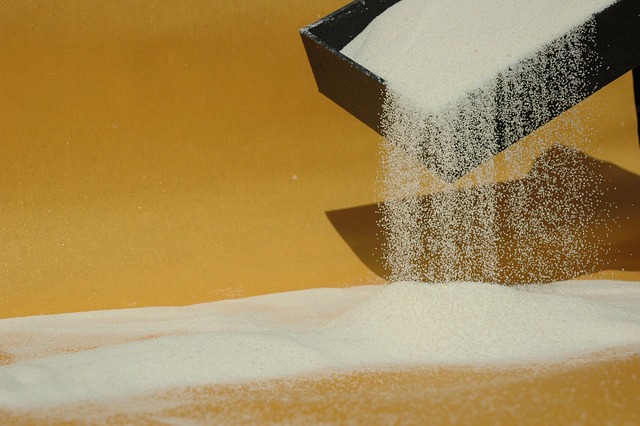
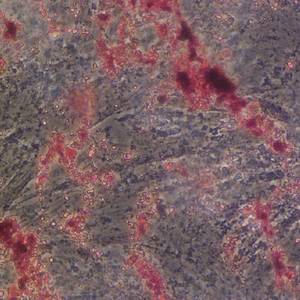
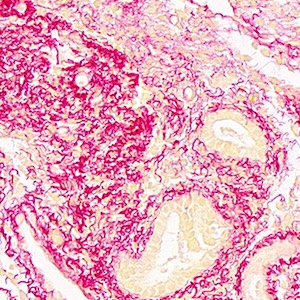
Morphological analysis of the seeds of three pseudocereals by using light microscopy and ESEM-EDS
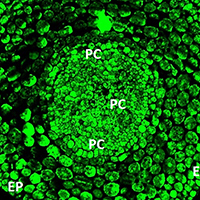
Submitted: 12 October 2019
Accepted: 30 December 2019
Published: 10 January 2020
Accepted: 30 December 2019
Abstract Views: 1419
PDF: 1027
HTML: 60
HTML: 60
Publisher's note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Similar Articles
- A.C. Croce, G. Bottiroli, Autofluorescence spectroscopy and imaging: a tool for biomedical research and diagnosis , European Journal of Histochemistry: Vol. 58 No. 4 (2014)
- Y. Kaneko, N. Onda, Y. Watanabe, M. Shibutani, Identification of 5-hydroxytryptamine-producing cells by detection of fluorescence in paraffin-embedded tissue sections , European Journal of Histochemistry: Vol. 60 No. 3 (2016)
- B. Sainz Jr., I. Miranda-Lorenzo, C. Heeschen, The fuss over lipo“fussâ€cin: not all autofluorescence is the same , European Journal of Histochemistry: Vol. 59 No. 1 (2015)
- F. Frontalini, D. Curzi, F.M. Giordano, J.M. Bernhard, E. Falcieri, R. Coccioni, Effects of lead pollution on Ammonia parkinsoniana (foraminifera): ultrastructural and microanalytical approaches , European Journal of Histochemistry: Vol. 59 No. 1 (2015)
- Anett Kristin Larsen, Jaione Simón-Santamaría, Kjetil Elvevold, Bo Göran Ericzon, Kim Erlend Mortensen, Peter McCourt, Bård Smedsrød, Karen Kristine Sørensen, Autofluorescence in freshly isolated adult human liver sinusoidal cells , European Journal of Histochemistry: Vol. 65 No. 4 (2021)
- D. Curzi, D. Lattanzi, S. Ciuffoli, S. Burattini, R.E. Grindeland, V.R. Edgerton, R.R. Roy, J.G. Tidball, E. Falcieri, Growth hormone plus resistance exercise attenuate structural changes in rat myotendinous junctions resulting from chronic unloading , European Journal of Histochemistry: Vol. 57 No. 4 (2013)
- Carlo Dall'Oca, Tommaso Maluta, Gian Mario Micheloni, Matteo Cengarle, Giampaolo Morbioli, Paolo Bernardi, Andrea Sbarbati, Daniele Degl'Innocenti, Franco Lavini, Bruno Magnan, The biocompatibility of bone cements: progress in methodological approach , European Journal of Histochemistry: Vol. 61 No. 2 (2017)
- Zhao Zhang, Hongming Fan, William Richardson, Bruce Z. Gao, Tong Ye, Management of autofluorescence in formaldehyde-fixed myocardium: choosing the right treatment , European Journal of Histochemistry: Vol. 67 No. 4 (2023)
- Francesca Scolari, Alessandro Girella, Anna Cleta Croce, Imaging and spectral analysis of autofluorescence patterns in larval head structures of mosquito vectors , European Journal of Histochemistry: Vol. 66 No. 4 (2022)
- E. Varricchio, M.G. Russolillo, L. Maruccio, S. Velotto, G. Campanile, M. Paolucci, F. Russo, Immunological detection of m- and µ-calpains in the skeletal muscle of Marchigiana cattle , European Journal of Histochemistry: Vol. 57 No. 1 (2013)
You may also start an advanced similarity search for this article.

 https://doi.org/10.4081/ejh.2020.3075
https://doi.org/10.4081/ejh.2020.3075