Smart Citations
Smart CitationsSee how this article has been cited at scite.ai
scite shows how a scientific paper has been cited by providing the context of the citation, a classification describing whether it supports, mentions, or contrasts the cited claim, and a label indicating in which section the citation was made.
Expression, localization and synthesis of small leucine-rich proteoglycans in developing mouse molar tooth germ
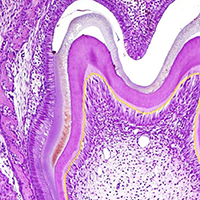
The gene expression and protein synthesis of small leucine-rich proteoglycans (SLRPs), including decorin, biglycan, fibromodulin, and lumican, was analyzed in the context of the hypothesis that they are closely related to tooth formation. In situ hybridization, immunohistochemistry, and organ culture with metabolic labeling of [35S] were carried out in mouse first molar tooth germs of different developmental stages using ICR mice at embryonic day (E) 13.5 to postnatal day (P) 7.0. At the bud and cap stage, decorin mRNA was expressed only in the surrounding mesenchyme, but not within the tooth germ. Biglycan mRNA was then expressed in the condensing mesenchyme and the dental papilla of the tooth germ. At the apposition stage (late bell stage), both decorin and biglycan mRNA were expressed in odontoblasts, resulting in a switch of the pattern of expression within the different stages of odontoblast differentiation. Decorin mRNA was expressed earlier in newly differentiating odontoblasts than biglycan. With odontoblast maturation and dentin formation, decorin mRNA expression was diminished and localized to the newly differentiating odontoblasts at the cervical region. Simultaneously, biglycan mRNA took over and extended its expression throughout the new and mature odontoblasts. Both mRNAs were expressed in the dental pulp underlying the respective odontoblasts. At P7.0, both mRNAs were weakly expressed but maintained their spatial expression patterns. Immunostaining showed that biglycan was localized in the dental papillae and pulp. In addition, all four SLRPs showed clear immunostaining in predentin, although the expressions of fibromodulin and lumican mRNAs were not identified in the tooth germs examined. The organ culture data obtained supported the histological findings that biglycan is more predominant than decorin at the apposition stage. These results were used to identify biglycan as the principal molecule among the SLRPs investigated. Our findings indicate that decorin and biglycan show spatial and temporal differential expressions and play their own tissue-specific roles in tooth development.
Downloads
Publication Facts
Reviewer profiles N/A
Author statements
- Academic society
- N/A
- Publisher
- PAGEPress Publications, Pavia, Italy
Citations
10.1007/s12565-020-00588-2
10.1093/glycob/cwae024
10.1007/s12565-022-00647-w
10.1007/s00441-022-03575-3
10.1093/g3journal/jkaf003
10.1186/s12864-023-09140-8
10.1177/00220345231224228
10.1007/s00441-024-03886-7
10.1007/978-3-030-76283-4_6
10.1016/j.jgg.2023.02.008
10.1038/s41368-023-00218-3
How to Cite
PAGEPress has chosen to apply the Creative Commons Attribution NonCommercial 4.0 International License (CC BY-NC 4.0) to all manuscripts to be published.

 https://doi.org/10.4081/ejh.2020.3092
https://doi.org/10.4081/ejh.2020.3092