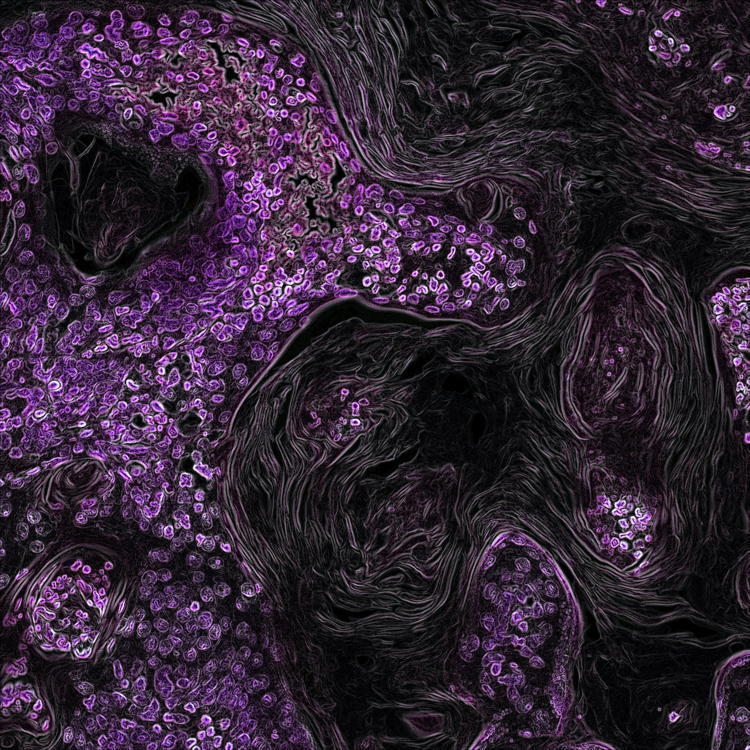
KISS1 and KISS1R expression in the human and rat carotid body and superior cervical ganglion
Submitted: 23 January 2011
Accepted: 22 March 2011
Published: 4 May 2011
Accepted: 22 March 2011
Abstract Views: 749
PDF: 417
HTML: 1009
HTML: 1009
Publisher's note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Similar Articles
- J.P. Damico, E. Ervolino, K.R. Torres, D.S. Batagello, R.J. Cruz-Rizzolo, C.A. Casatti, J.A. Bauer, Phenotypic alterations of neuropeptide Y and calcitonin gene-related peptide-containing neurons innervating the rat temporomandibular joint during carrageenan-induced arthritis , European Journal of Histochemistry: Vol. 56 No. 3 (2012)
- Ying Wang, Shifa Yuan, Jing Ma, Hong Liu, Lizhen Huang, Fengzhen Zhang, Substance P is overexpressed in cervical squamous cell carcinoma and promoted proliferation and invasion of cervical cancer cells in vitro , European Journal of Histochemistry: Vol. 67 No. 3 (2023)
- A.K. Lindström, D. Hellberg, Immunohistochemical LRIG3 expression in cervical intraepithelial neoplasia and invasive squamous cell cervical cancer: association with expression of tumor markers, hormones, high-risk HPV-infection, smoking and patient outcome , European Journal of Histochemistry: Vol. 58 No. 2 (2014)
- Liyun Guan, Shifa Yuan, Jing Ma, Hong Liu, Lizhen Huang, Fengzhen Zhang, Neurokinin-1 receptor is highly expressed in cervical cancer and its antagonist induces cervical cancer cell apoptosis , European Journal of Histochemistry: Vol. 67 No. 1 (2023)
- L. Ragionieri, M. Botti, F. Gazza, C. Sorteni, R. Chiocchetti, P. Clavenzani, L. Bo, R. Panu, Localization of peripheral autonomic neurons innervating the boar urinary bladder trigone and neurochemical features of the sympathetic component , European Journal of Histochemistry: Vol. 57 No. 2 (2013)
- R. Pennati, A. Dell'Anna, G. Zega, F. De Bernardi, Immunohistochemical study of the nervous system of the tunicate Thalia democratica (Forsskal, 1775) , European Journal of Histochemistry: Vol. 56 No. 2 (2012)
- J. Melrose, The knee joint loose body as a source of viable autologous human chondrocytes , European Journal of Histochemistry: Vol. 60 No. 2 (2016)
- W. Meyer, M. Liumsiricharoen, A. Suprasert, L.G. Fleischer, M. Hewicker-Trautwein, Immunohistochemical demonstration of keratins in the epidermal layers of the Malayan pangolin (Manis javanica), with remarks on the evolution of the integumental scale armour , European Journal of Histochemistry: Vol. 57 No. 3 (2013)
- G. Scarsella, M. Nebbioso, S. Stefanini, N. Pescosolido, Degenerative effects in rat eyes after experimental ocular hypertension , European Journal of Histochemistry: Vol. 56 No. 4 (2012)
- Wenjing Liu, Shanshan Ming, Xiaobing Zhao, Xin Zhu, Yuxiang Gong, Developmental expression of high-mobility group box 1 (HMGB1) in the mouse cochlea , European Journal of Histochemistry: Vol. 67 No. 3 (2023)
You may also start an advanced similarity search for this article.

 https://doi.org/10.4081/ejh.2011.e14
https://doi.org/10.4081/ejh.2011.e14